What's Cooking - Multiclass Imbalance
This post is a continuation of my series of posts on the now extremely expired Kaggle competition What’s Cooking. In my previous post, Hello World Model, I created a basic random forest model in R and found that while its overall accuracy was quite high (93%), its balanced accuracy (76%) could be improved. In this post, I’ll look at some strategies for accommodating class imbalance while still using the same random forest multiclass classifier.
There are several potential options built into the randomForest() package that can help us right off. The first I’ll explore is the classwt parameter, from the documentation, this parameter sets the ‘priors of the classes’ and ‘it need not add to 1’. By default, all classes have a classwt = 1, meaning that a misclassification error during training counts the same for all classes. By changing this parameter so that one class has a classwt of 2 and the rest have a classwt of 1, you are telling the RandomForest that a misclassification is double penalized for one class versus another. This forces the model to minimize error in the class with higher weight. We can supply any numeric vector that is equal to the number of classes to R and not worry about it adding to 1 because when the model is compiled, the classwts are regularized. I tested two strategies for class weights. The first is a basic inverse frequency weighting:
cuisine_freq = table(train_test$train_dt_y)
base_wts = 1/(cuisine_freq/min(cuisine_freq))
rfwts_1 = randomForest(x = train_test$train_dt_x,
y = train_test$train_dt_y,
ntree = 1500,
mtry = 10,
classwt = base_wts)Here the smallest class (british) has a classwt = 1 and the largest class (mexican) has a classwt of .03. This model resulted in a slightly weaker overall accuracy (weighted : 88%, base : 93%) but a much stronger balanced accuracy (weighted : 90% , base : 76%). Looking at the plot below, we can see that the model gave strong preference to the smaller classes, causing large class performance to suffer.
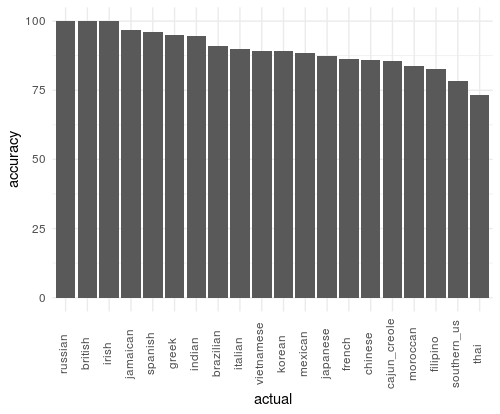
I tested using a slightly less drastic weighting method to see if I could get better class accuracy balance:
cuisine_freq = table(train_test$train_dt_y)
new_wts = log(cuisine_freq/min(cuisine_freq))
new_wts[is.infinite(new_wts)] = 1
new_wts[new_wts < 1] = 1
new_wts = 1/new_wts
rf_wts_2 = randomForest(x = train_test$train_dt_x,
y = train_test$train_dt_y,
ntree = 2500,
mtry = 10,
classwt = new_wts)
Which improved overall accuracy (92%) and balanced accuracy (92%)
Another option built into the randomForest() package is to use stratified sampling in the creation of each tree. By default, the model will sample your data, with replacement, to create a bootstrapped dataset for each tree it constructs. Each observation in your data is equally likely to occur in this bootstrapped dataset, so, naturally, larger classes have larger representation. We can alter this behavior by telling the model that we want to take a specific number of samples from each class, using the sampsize agrument. By giving a set number from each class to sample, we can make each tree have equal representation for all classes. NOTE - it is necessary to set replace = FALSE when doing this to keep your smaller classes from having overly influential observations. Like the classwt parameter, I tried two different configurations of the sampsize parameter to improve performance.
First, I tried setting the samp size to be equal across all classes. To do this, I took 80% of the smallest class size and propagated this sample size to all classes. This resulted in poorer performance than weighting but better than the base model, (overall accuracy : 91%, balanced accuracy : 85%).
cuisine_freq = table(train_test$train_dt_y)
samp_prop = .8
equal_samp_size = round(min(cuisine_freq)*samp_prop)
samp_vec = rep(equal_samp_size, length(cuisine_freq))
names(samp_vec) = names(cuisine_freq)
rf_samp_1 = randomForest(x = train_test$train_dt_x,
y = train_test$train_dt_y,
ntree = 2500,
mtry = 10,
sampsize = samp_vec,
replace = F)
Thinking I could get a similar improvement to accuracy by making the samp size a bit more representative of the data, I tried another sample size method :
cuisine_freq = table(train_test$train_dt_y)
samp_prop = .8
equal_samp_size = round(cuisine_freq*samp_prop)
new_sample_prop = log(equal_samp_size/min(equal_samp_size))
new_sample_prop[new_sample_prop<1]= 1
new_samp_vec = round(new_sample_prop*min(equal_samp_size))
rf_samp_2 = randomForest(x = train_test$train_dt_x,
y = train_test$train_dt_y,
ntree = 2500,
mtry = 10,
sampsize = new_samp_vec,
replace = F)Unfortunately, it did not improve performance - in fact its performance is very similar to the base model (overall accuracy : 93%, balanced accuracy : 79%).
There are many other ways we could address the class imbalance outside of model configurations - these change the training data by either over sampling the smaller classes, or undersampling the larger ones, or create new artificial minority class examples via SMOTE. I won’t cover these here but they may be useful in the future.
As this post shows, setting these optional parameters can really impact the performance of your model. It also shows that it can quickly get overwhelming keeping track of all the different configurations and resulting performance data. In my next post, I’ll talk about how we can deploy hyperparameter tuning to make this process more automated.
